Converts a genlight object into a format suitable for input to the BPP program
Source:R/gl2bpp.r
gl2bpp.RdThis function generates the sequence alignment file and the Imap file. The control file should produced by the user.
gl2bpp(
x,
method = 1,
outfile = "output_bpp.txt",
imap = "Imap.txt",
outpath = tempdir(),
verbose = NULL
)Arguments
- x
Name of the genlight object containing the SNP data [required].
- method
One of 1 | 2, see details [default = 1].
- outfile
Name of the sequence alignment file ["output_bpp.txt"].
- imap
Name of the Imap file ["Imap.txt"].
- outpath
Path where to save the output file (set to tempdir by default)
- verbose
Verbosity: 0, silent or fatal errors; 1, begin and end; 2, progress log; 3, progress and results summary; 5, full report [default 2 or as specified using gl.set.verbosity].
Value
returns no value (i.e. NULL)
Details
If method = 1, heterozygous positions are replaced by standard ambiguity codes.
If method = 2, the heterozygous state is resolved by randomly assigning one or the other SNP variant to the individual.
Trimmed sequences for which the SNP has been trimmed out, rarely, by adapter mis-identity are deleted.
This function requires 'TrimmedSequence' to be among the locus metrics
(@other$loc.metrics) and information of the type of alleles (slot
loc.all e.g. 'G/A') and the position of the SNP in slot position of the
“`genlight“` object (see testset.gl@position and testset.gl@loc.all for
how to format these slots.)
It's important to keep in mind that analyses based on coalescent theory, like those done by the programme BPP, are meant to be used with sequence data. In this type of data, large chunks of DNA are sequenced, so when we find polymorphic sites along the sequence, we know they are all on the same chromosome. This kind of data, in which we know which chromosome each allele comes from, is called "phased data." Most data from reduced representation genome-sequencing methods, like DArTseq, is unphased, which means that we don't know which chromosome each allele comes from. So, if we apply coalescence theory to data that is not phased, we will get biased results. As in Ellegren et al., one way to deal with this is to "haplodize" each genotype by randomly choosing one allele from heterozygous genotypes (2012) by using method = 2.
Be mindful that there is little information in the literature on the validity of this method.
References
Ellegren, Hans, et al. "The genomic landscape of species divergence in Ficedula flycatchers." Nature 491.7426 (2012): 756-760.
Flouri T., Jiao X., Rannala B., Yang Z. (2018) Species Tree Inference with BPP using Genomic Sequences and the Multispecies Coalescent. Molecular Biology and Evolution, 35(10):2585-2593. doi:10.1093/molbev/msy147
Examples
require(dartR.data)
test <- platypus.gl
test <- gl.filter.callrate(test,threshold = 1)
#> Starting gl.filter.callrate
#> Processing genlight object with SNP data
#> Warning: data include loci that are scored NA across all individuals.
#> Consider filtering using gl <- gl.filter.allna(gl)
#> Warning: Data may include monomorphic loci in call rate
#> calculations for filtering
#> Recalculating Call Rate
#> Removing loci based on Call Rate, threshold = 1
#>
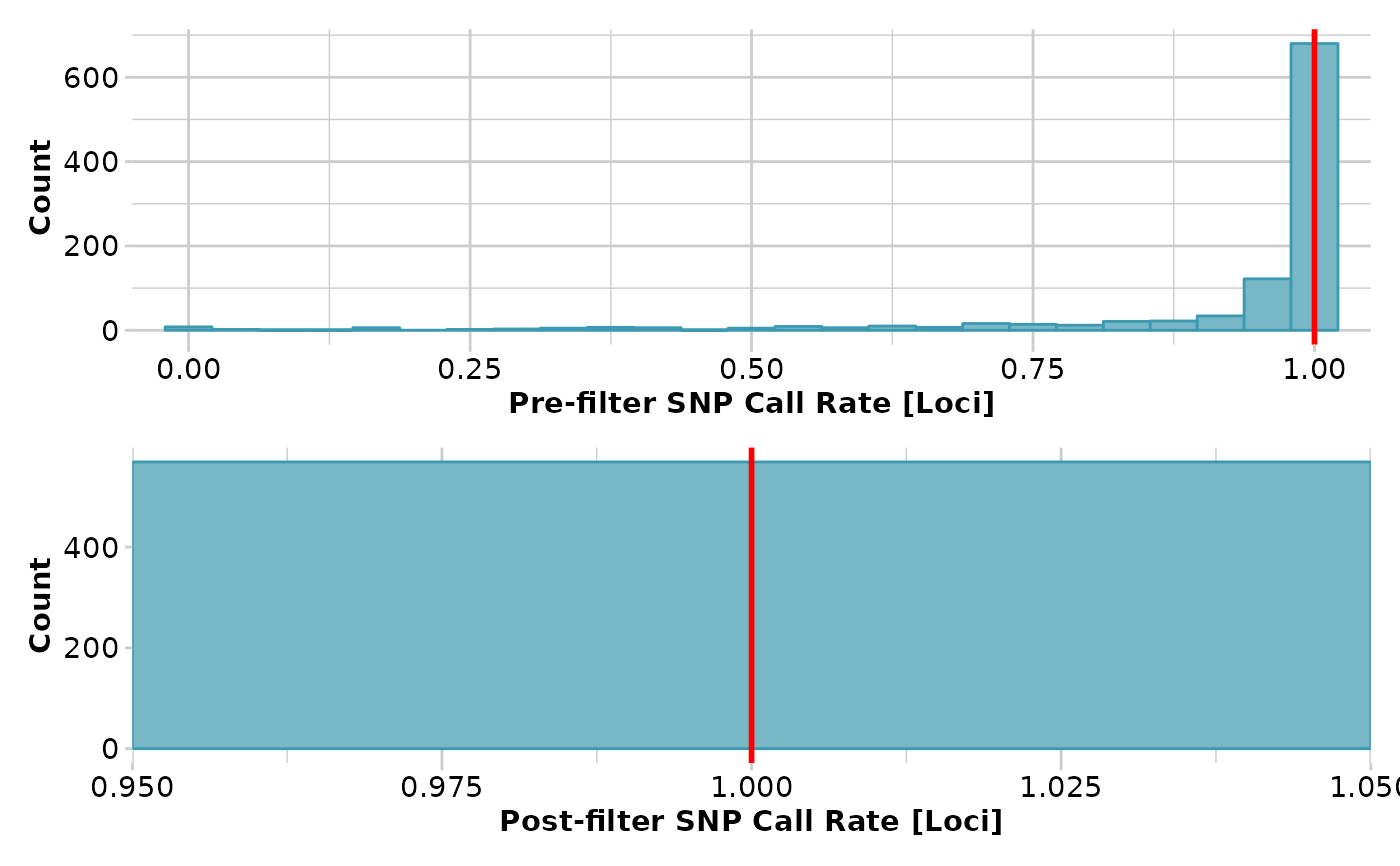 #> Completed: gl.filter.callrate
#>
test <- gl.filter.monomorphs(test)
#> Starting gl.filter.monomorphs
#> Processing genlight object with SNP data
#> Identifying monomorphic loci
#> Removing monomorphic loci and loci with all missing
#> data
#> Completed: gl.filter.monomorphs
#>
test <- gl.subsample.loci(test,n=25)
#> Starting gl.subsample.loci
#> Processing genlight object with SNP data
#> Subsampling at random 25 loci from dartR object
#> Completed: gl.subsample.loci
#>
gl2bpp(x = test)
#> Starting gl2bpp
#> Processing genlight object with SNP data
#> Assigning ambiguity codes to heterozygote SNPs
#> Removing loci for which SNP position is outside the length of the trimmed sequences
#> Generating haplotypes ... This may take some time
#> Completed: gl2bpp
#>
#> NULL
#> Completed: gl.filter.callrate
#>
test <- gl.filter.monomorphs(test)
#> Starting gl.filter.monomorphs
#> Processing genlight object with SNP data
#> Identifying monomorphic loci
#> Removing monomorphic loci and loci with all missing
#> data
#> Completed: gl.filter.monomorphs
#>
test <- gl.subsample.loci(test,n=25)
#> Starting gl.subsample.loci
#> Processing genlight object with SNP data
#> Subsampling at random 25 loci from dartR object
#> Completed: gl.subsample.loci
#>
gl2bpp(x = test)
#> Starting gl2bpp
#> Processing genlight object with SNP data
#> Assigning ambiguity codes to heterozygote SNPs
#> Removing loci for which SNP position is outside the length of the trimmed sequences
#> Generating haplotypes ... This may take some time
#> Completed: gl2bpp
#>
#> NULL